Behaviour and Ecology
@ School of Life Sciences,
University of Nottingham
| Home | People | Research | Teaching | Publications | Opportunities | Links |
Parasites and evolution - Andrew MacColl
Ecology has
been very successful at exploring certain types of interactions between
species
within natural systems, while generally ignoring the possibility that
such
interactions result in natural selection and can evolve. Evolutionary
biologists have been very good at demonstrating natural selection and
exploring
the genetic basis of selected phenotypes, but have tended to do so in
isolation
from their ecological settings. In the past decade there has been a resurgence
of interest in reconnecting these closely related but disjointed
perspectives,
with a view to better understanding how mechanism (ecology) and process
(natural selection) lead to pattern (evolution). The main theme of Andrew MacColl's research
is to understand the role of ecology in driving natural selection and
how this
can produce the divergent evolution that builds into variation between
individuals,
populations and that ultimately accumulates into speciation.
understanding how mechanism (ecology) and process
(natural selection) lead to pattern (evolution). The main theme of Andrew MacColl's research
is to understand the role of ecology in driving natural selection and
how this
can produce the divergent evolution that builds into variation between
individuals,
populations and that ultimately accumulates into speciation.
A Schistocephalus solidus tapeworm dissected out of the body cavity of a dead stickleback. These worms can be almost the same weight as their host!
Our
approach is based on the integration of theory, observation and
experiment. We are particularly interested in the relative importance of different
selective
agents in directing evolution. In general little is known about whether
evolution is driven mainly by the abiotic environment or by ecological
interactions such as competition, predation and parasitism. Classical
ecology
has focussed on the part played by competition between organisms in
determining
individual success (and hence evolution). Other ecological interactions
have
received less attention. At present we are particularly interested in
the role
of parasitism as a driver of host evolution, because parasitism has
been poorly
studied from this perspective.
Stickleback as a model for the study of evolution
Three-spined
sticklebacks (Gasterosteus aculeatus)
are a good model species because they are common, widely distributed
and easy
to keep in the lab. A great deal is known about their natural history
and
genetics (their genome has been sequenced). They are particularly
interesting
because they exhibit a great deal of phenotypic and genetic diversity
between
populations. The photographs below show sticklebacks from different
populations
on the
Figure
1.
Morphological variation between five populations of three-spined
sticklebacks
on North Uist in the
We use
three-spined sticklebacks to address the following kinds of general
questions:
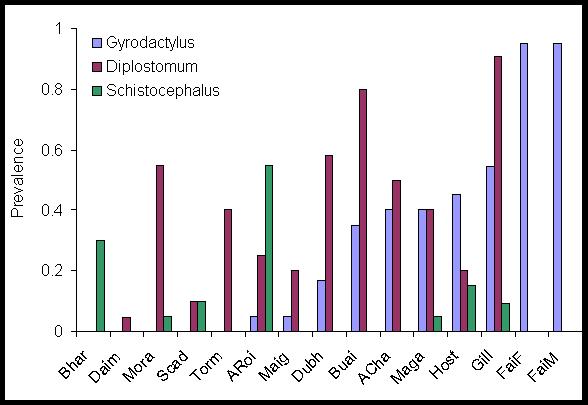
Figure 2. Variation in the prevalence of the three commonest macroparasites of sticklebacks (N=20 per loch) in 15 lochs in a 1000 km2 area of North Uist.
This graph clearly shows that
there are large differences in parasitism between
host populations even in small geographical areas. Initial results show
that
these differences are fairly stable across years.
Is the change in selection caused by parasites important when hosts invade novel environments?
This is equivalent to asking whether
hosts become adapted to the
parasites in their own population. One way to
look at this is to simulate invasions of novel environments, by
carrying out
transplant experiments. We have introduced lab raised fish into
enclosures in
lochs and monitored their growth and parasite burdens after a month.
The
parents of these fish came either from that loch (sympatric) or from a
different loch (allopatric). Some results are shown in Figure 3.
3
Introducing sticklebacks into enclosures in a freshwater loch on North Uist.
3
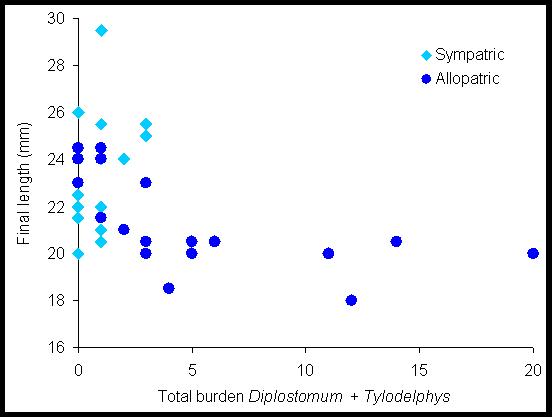
Figure 3. The relationship between the burden of trematode parasites in the eyes of fish and their length at the end of the experiment. Fish were 17-18mm when the experiment began. The figure shows two things: (i) Allopatric sticklebacks suffer higher parasite burdens than sympatric ones. (ii) The growth of infected allopatric fish is reduced compared to uninfected allopatric fish or to sympatric fish. Taken together these suggest that parasites can be an important agent of selection on hosts.
The data in
Figure 3 suggest that parasites could cause selection on life history
traits.
This possibility has been discussed at length theoretically, but as yet
there
are few relevant data. There is certainly substantial variation in life
history
characters between stickleback populations, and this is associated with
variation in parasite prevalence (Fig. 4). Some of our current research
is
directed at determining whether such relationships have a causal basis.
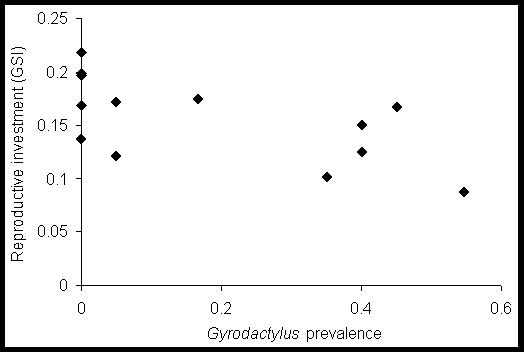
Figure
4.
The relationship between prevalence of Gyrodactylus
arcuatus and the
reproductive investment of female sticklebacks
(gonadosomatic index, GSI) across freshwater populations on North Uist
(P<0.025).
It is also
likely that variation in parasitism between populations should lead to
variation in the ability of hosts to resist or tolerate their
parasites. This
can be investigated by exposing lab raised sticklebacks from different
populations to infection with standard dose of parasites in the lab.
Limnetic
and benthic sticklebacks that occur together in lakes in British
Columbia
exhibit very different parasite loads in the wild and also have very
different
resistance to artificial infection with the parasite Diplostomum
scudderi (Fig. 5). Crosses between these two species
result in hybrids that have the same susceptibility to infection as the
limnetic grandparents, suggesting that susceptibility may be dominant
and
contribute to the poor performance of hybrids in the wild.
We are
beginning to explore differences in resistance traits between Scottish
populations of sticklebacks, and whether such differences are are
involved in
trade-offs with life history and mate choice traits.
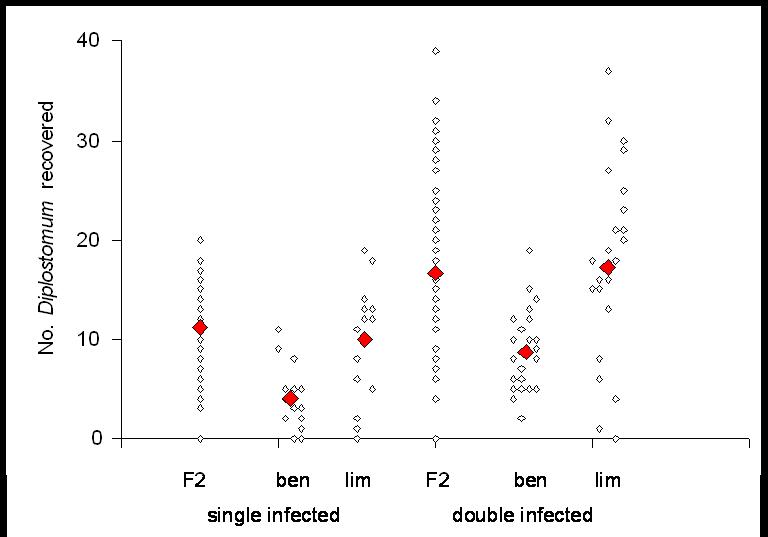
Figure
5.
Numbers of parasites recovered from sticklebacks artificially infected
with one
or two doses of twenty Diplostomum
scudderi.
‘ben’ = benthic, ‘lim’ =
limnetic and F2 = second generation
hybrids, all lab reared fish originally from Paxton lake,
Can the interaction between hosts and parasites lead to divergence between host populations that results in reproductive isolation between those populations?
The results shown in figure 5 (above) suggest that hybrids between divergent
host populations may have poor resistance to parasites. This is a good example
of how parasites could contribute to ecologically determined post-mating
reproductive isolation. We
are beginning to explore other ways in which parasite imposed selection
could
contribute to accumulation of reproductive isolation between host
populations.
What next?
We are also
beginning to explore whether the traits of parasites and hosts coevolve
in our
study systems, including such questions as:
Does the
selection that parasites impose on their hosts result in coevolutionary
feedback to the parasites, because of changes in host density or mean
host
phenotypes?
Does
coevolutionary dynamism result in arms races between parasites and
hosts that
can lead in fundamentally unpredictable directions?